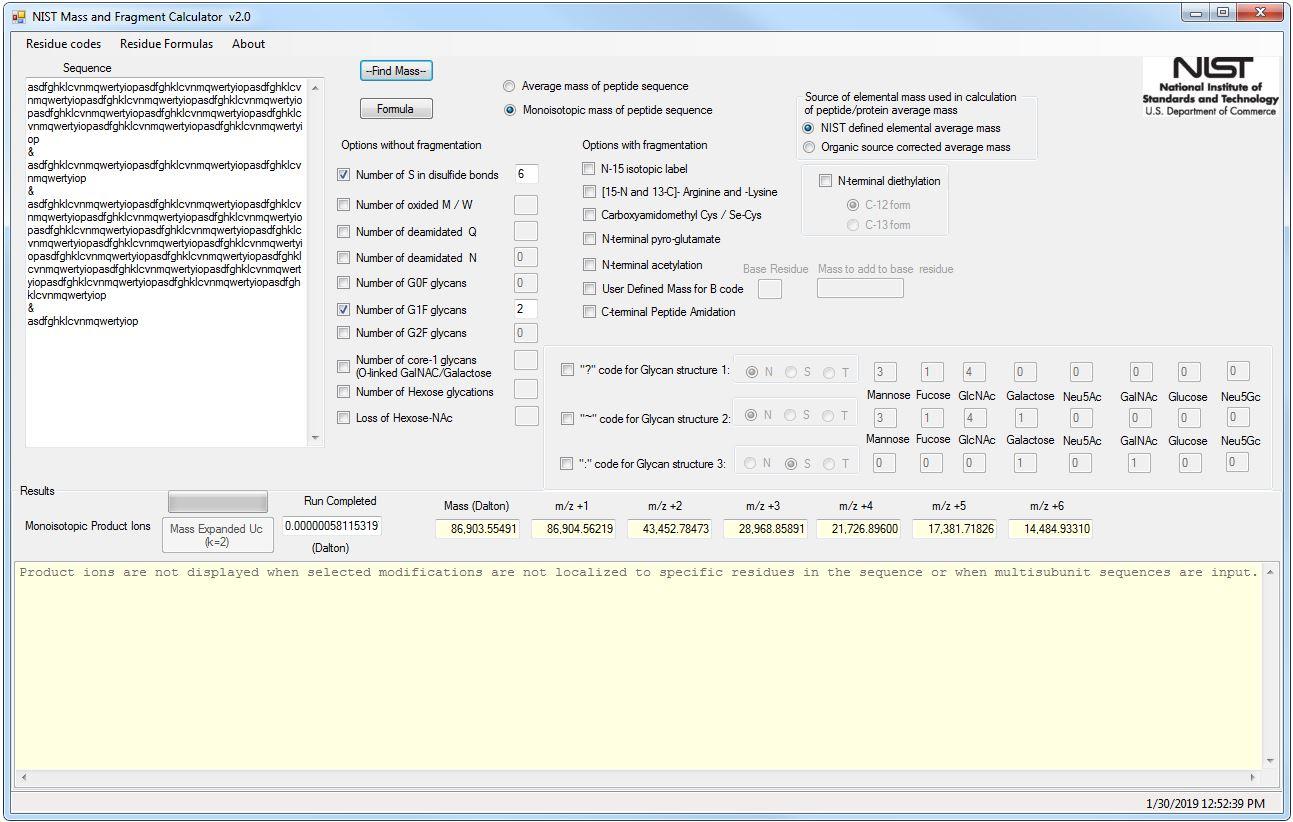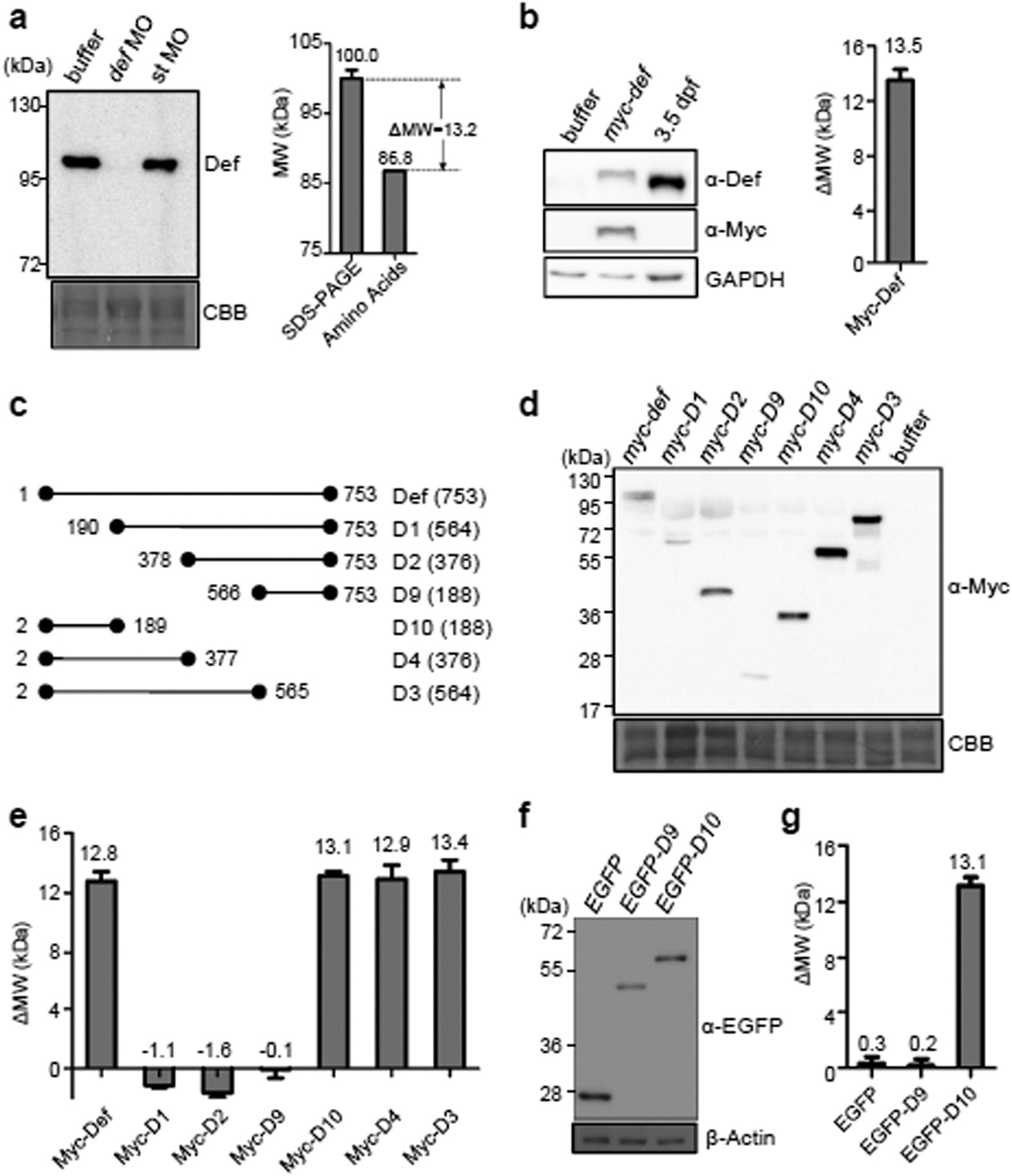
An equation to estimate the difference between theoretically predicted and SDS PAGE-displayed molecular weights for an acidic peptide | Scientific Reports

PDBcor: An automated correlation extraction calculator for multi-state protein structures - ScienceDirect
Accurate calculation of side chain packing and free energy with applications to protein molecular dynamics | PLOS Computational Biology
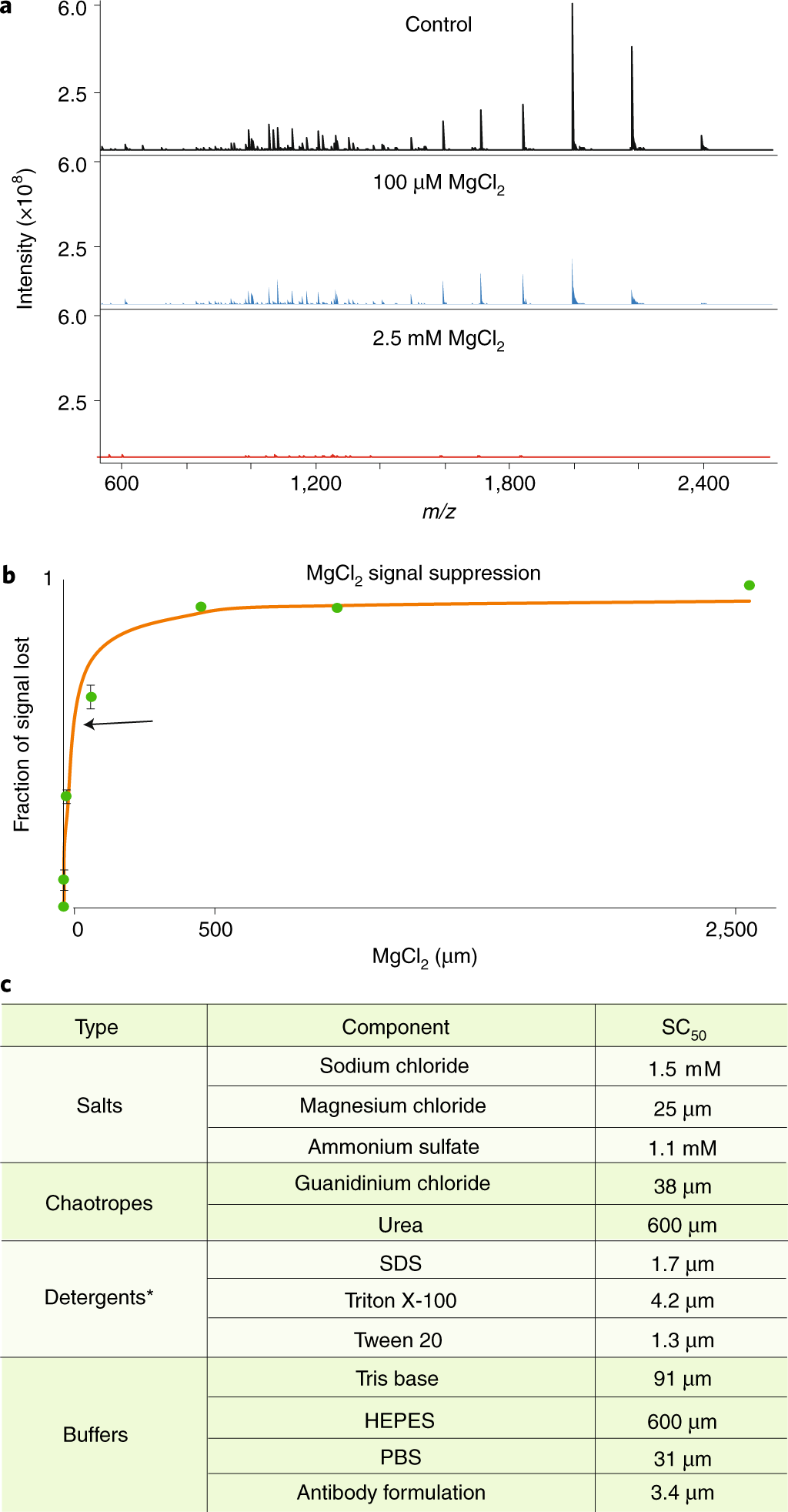
Best practices and benchmarks for intact protein analysis for top-down mass spectrometry | Nature Methods
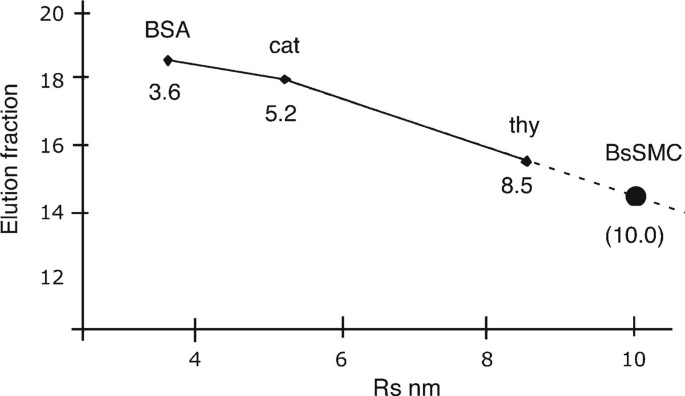
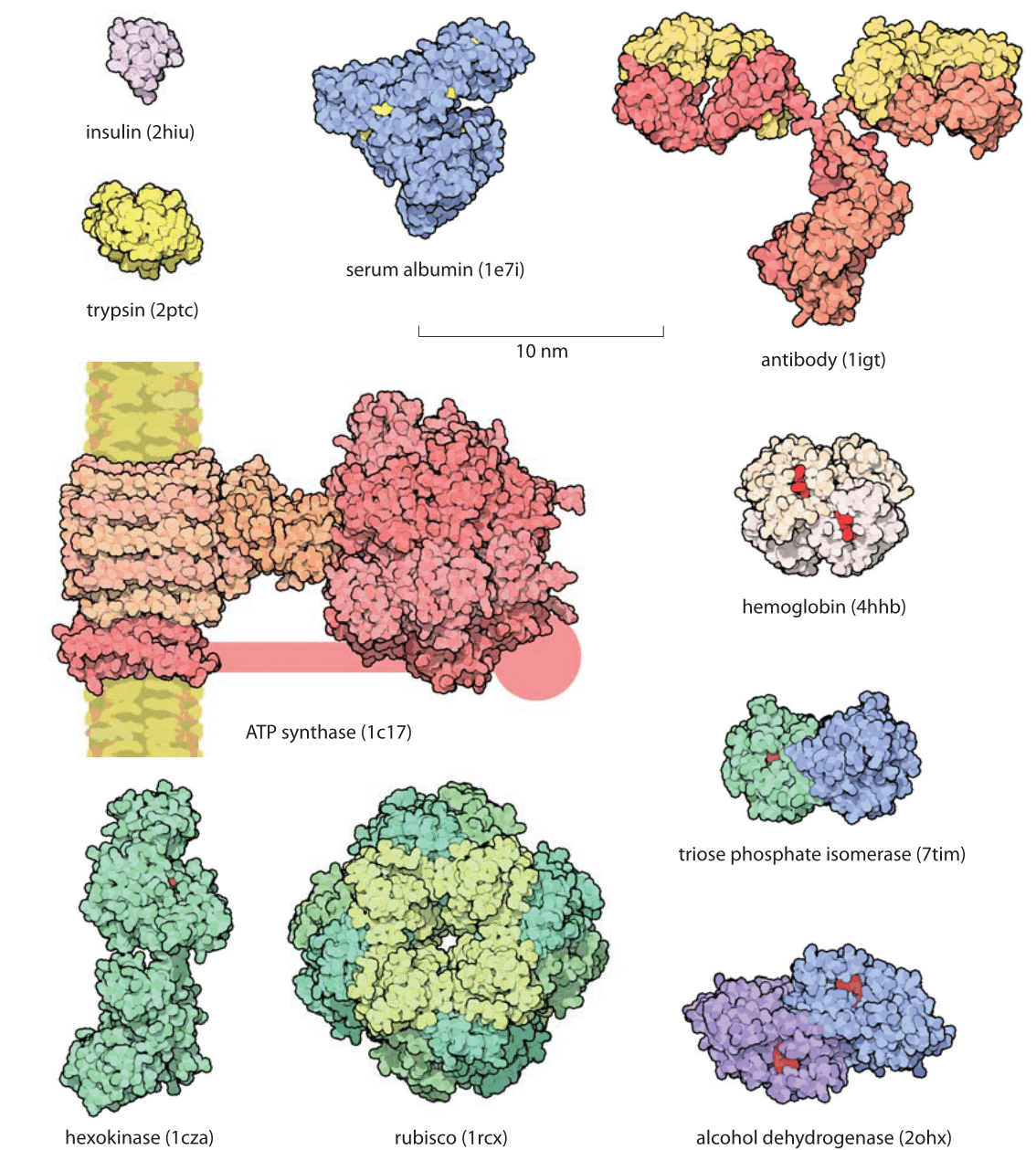
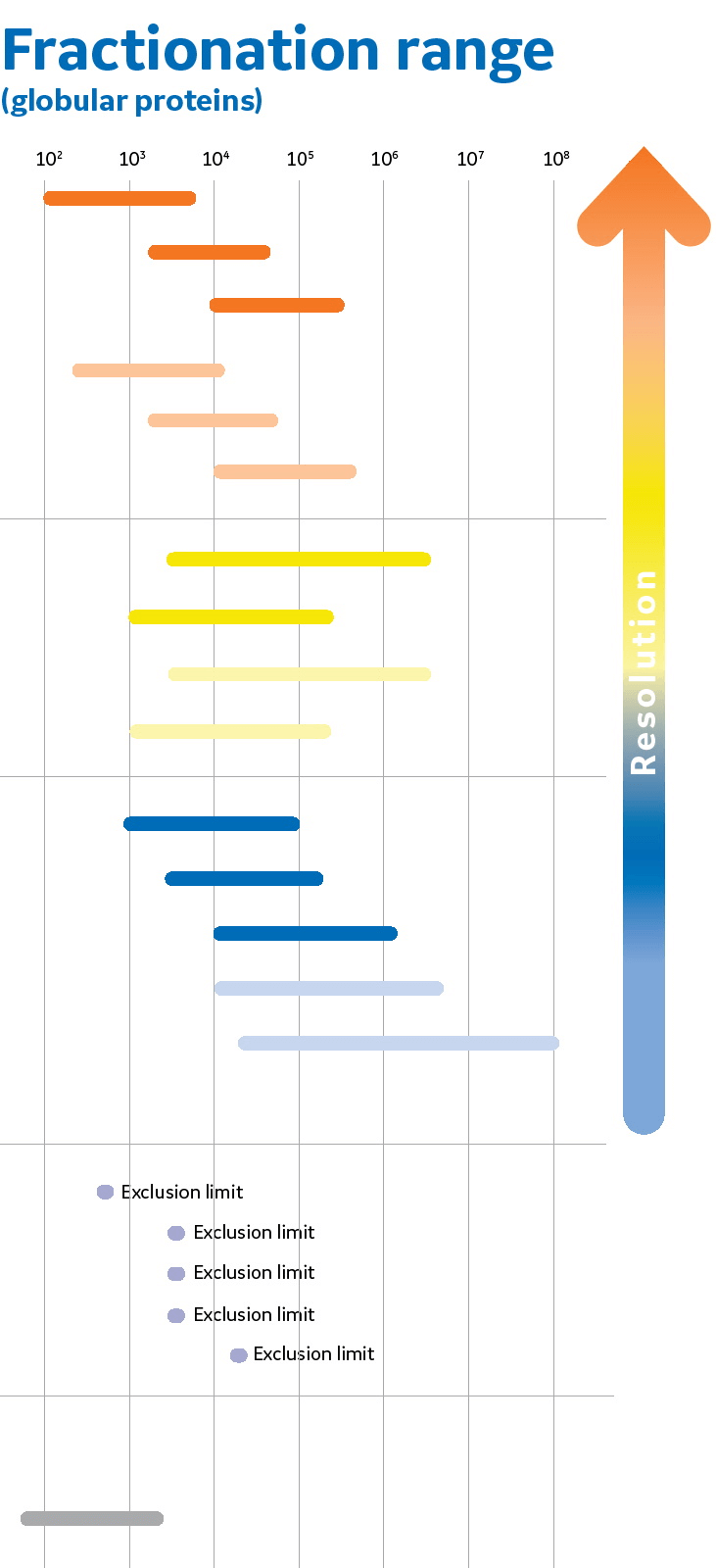
Size and Shape of Protein Molecules at the Nanometer Level Determined by Sedimentation, Gel Filtration, and Electron Microscopy | Biological Procedures Online | Full Text

SAXSMoW 2.0: Online calculator of the molecular weight of proteins in dilute solution from experimental SAXS data measured on a relative scale - Piiadov - 2019 - Protein Science - Wiley Online Library